 Hahsler, M., Hornik, K.: TSP-infrastructure for the traveling salesperson problem. Hahsler, M., Hornik, K., Buchta, C.: Getting things in order: an introduction to the R package seriation. Liiv, I.: Seriation and matrix reordering methods: an historical overview. Kendall, D.G.: Abundance matrices and seriation in archaeology. 2(4), 317–334 (1995)įulkerson, D., Gross, O.: Incidence matrices and interval graphs. 28(1), 197–226 (2011)īarnard, S.T., Pothen, A., Simon, H.: A spectral algorithm for envelope reduction of sparse matrices. Garriga, G.C., Junttila, E., Mannila, H.: Banded structure in binary matrices. 
Robinson, W.S.: A method for chronologically ordering archaeological deposits. Hahsler, M.: An experimental comparison of seriation methods for one-mode two-way data. It is showed that the proposed method outperforms the other state-of-the-art methods used for comparison purposes in this specific type of data. Experimental results based on both synthetic and real data are used to validate and illustrate the application of the method. This similarity information is based on Histogram Oriented Gradient (HOG) features, computed from random locations at the images in each iteration/revisitation. Instead, we propose a graph-based method where the images are iteratively revisited and the image similarity information is refined, until a linear graph representing the seriation is obtained. However, these methods are very sensitive to noise, distortion, and missing data, since the optimal solution for the matrix does not match the true seriation. Most image seriation methods are two-step: firstly, a distance matrix is obtained from image processing, and secondly, the optimal seriation method is determined for this matrix. It is also a challenging problem since, during the manual procedure of cutting the brains and acquiring hundreds of images, the human operator can unwittingly change its natural sequence, loose data, induce large morphological distortions, or introduce artifacts. This is fundamental for the neuroscience community to help in the processing and analyzing the huge amount of experimental data. 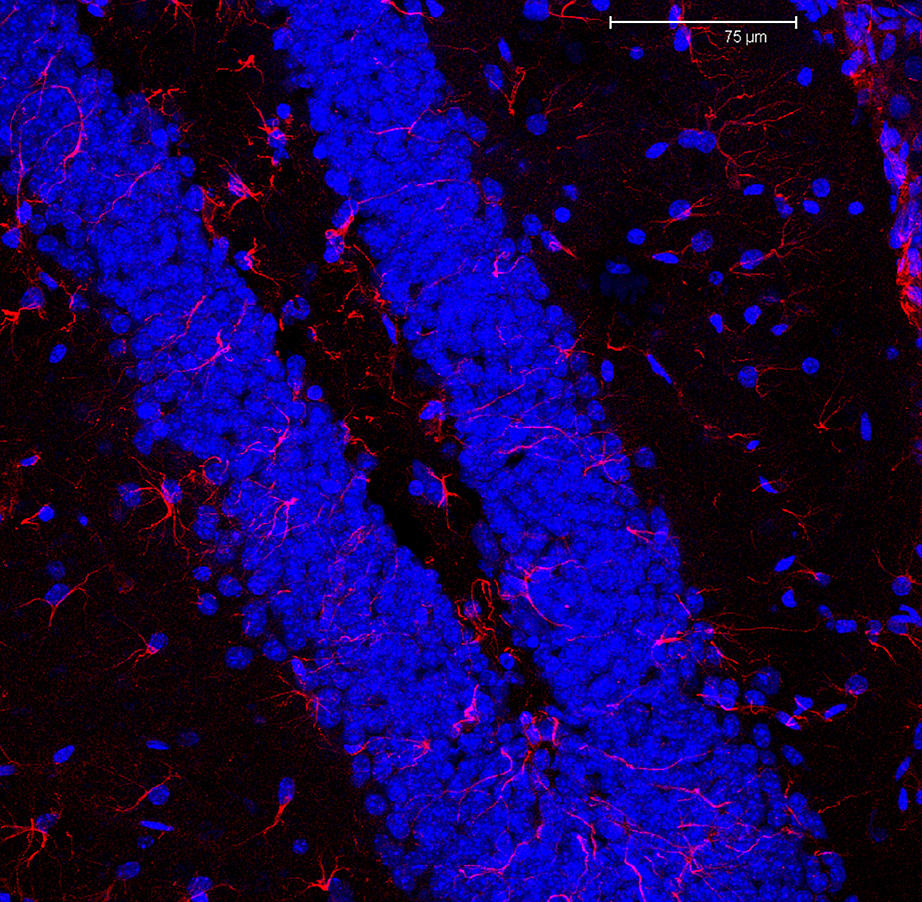
All rights reserved.This paper addresses the problem of automatic seriation of mouse brain cross-sections stained with green-florescence protein (GFP). We propose that our automated brain mapping method enables greater efficiency and consistency in mouse neuropathologic assessments.Īutomated image analysis Brain atlas Brain map Image segmentation Mouse histologic sections.Ĭopyright © 2018 Elsevier B.V. Optical density quantification results were comparable to those from time-consuming, manually drawn hippocampal delineations on the IHC-stained sections.Ĭompared to other published methods, our method requires less manual input, and has been validated comprehensively using a secondary atlas, as well as manually annotated brain IHC sections from 68 study mice. Nissl-stained sections were mapped and hippocampal boundaries transferred to adjacent immunohistochemically stained sections. We subsequently applied our method to mouse brain sections from an in vivo study where the hippocampus was the structure of interest. Brain regions mapped from FP Nissl-stained sections and calculated volumes were similar to structure delineations and volumes derived from corresponding FP illustrations. 
The method's accuracy was first assessed by comparing the mapping results to structure delineations from the Franklin and Paxinos (FP) mouse brain atlas. The method utilizes the publicly available Allen Brain Atlas to map brain regions on digitized Nissl-stained sections. We developed an automated brain mapping method that identifies neuroanatomic structures in mouse histologic coronal brain sections. Automated image analysis is a powerful tool that can radically reduce the workload associated with evaluating brain histologic sections. Histologic evaluation of the central nervous system is often a critical endpoint in in vivo efficacy studies, and is considered the essential component of neurotoxicity assessment in safety studies. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
